Cellular Disulfide Bond Formation in Bioactive Peptides and Proteins
Abstract
:1. Introduction
2. Folding of Peptides and Proteins in Normal Cells

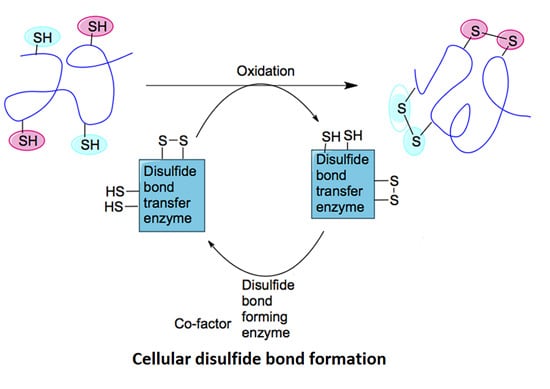
2.1. Mechanisms of Disulfide Bond Formation

| Site | Disulfide Bond Transferring Enzyme | Disulfide Bond Generating Enzyme | Cofactor |
|---|---|---|---|
| Prokaryotic | DsbA | DsbB | Ubiquinone |
| Periplasm | DsbC | DsbD | Ubiquinone |
| DsbG | DsbD | - | |
| Endoplasmic reticulum | PDI | Ero1 | FAD |
| PDI | Erv2 | FAD | |
| Mitochondria | Mia40 | Erv1 | FAD |
| Chloroplast | PSII | LTO1 | Phylloquinone or Hydroquinone |
| PSII | LQY1 | Zn (believed to be a cofactor) | |
| PSI and PSII | CYO1 | Zn (believed to be a cofactor) | |
| Extracellular space | QSOX | QSOX | FAD |
2.1.1. Periplasmic System
2.1.2. Endoplasmic Reticulum System
2.1.3. Mitochondria
2.1.4. Chloroplasts
2.1.5. Extracellular Space
3. Folding of Disulfide-Containing Bioactive Peptides and Proteins via Recombinant Technology
3.1. Co-Expressing Supporting Enzymes/Peptides
3.2. Chemical Assistance in Recombinant Folding
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Moroder, L.; Musiol, H.J.; Gotz, M.; Renner, C. Synthesis of single- and multiple-stranded cystine-rich peptides. Biopolymers 2005, 80, 85–97. [Google Scholar] [CrossRef] [PubMed]
- Góngora-Benítez, M.; Tulla-Puche, J.; Albericio, F. Multifaceted roles of disulfide bonds. peptides as therapeutics. Chem. Rev. 2013, 114, 901–926. [Google Scholar] [CrossRef] [PubMed]
- Marshall, E.; Costa, L.M.; Gutierrez-Marcos, J. Cysteine-rich peptides (CRPs) mediate diverse aspects of cell-cell communication in plant reproduction and development. J. Exp. Bot. 2011, 62, 1677–1686. [Google Scholar] [CrossRef] [PubMed]
- Haber, E.; Anfinsen, C.B. Regeneration of enzyme activity by air oxidation of reduced subtilisin-modified ribonuclease. J. Biol. Chem. 1961, 236, 422–424. [Google Scholar] [PubMed]
- Anfinsen, C.B. Principles that govern the folding of protein chains. Science 1973, 181, 223–230. [Google Scholar] [CrossRef] [PubMed]
- Heras, B.; Kurz, M.; Shouldice, S.R.; Martin, J.L. The name’s bond......disulfide bond. Curr. Opin. Struct. Biol. 2007, 17, 691–698. [Google Scholar] [CrossRef] [PubMed]
- Kadokura, H.; Beckwith, J. Mechanisms of oxidative protein folding in the bacterial cell envelope. Antioxid. Redox Signal. 2010, 13, 1231–1246. [Google Scholar] [CrossRef] [PubMed]
- Sato, Y.; Inaba, K. Disulfide bond formation network in the three biological kingdoms, bacteria, fungi and mammals. FEBS J. 2012, 279, 2262–2271. [Google Scholar] [CrossRef] [PubMed]
- Inaba, K. Structural basis of protein disulfide bond generation in the cell. Genes Cells 2010, 15, 935–943. [Google Scholar] [CrossRef] [PubMed]
- Hiniker, A.; Bardwell, J.C. Disulfide relay between and within proteins: the Ero1p structure. Trends Biochem. Sci. 2004, 29, 516–519. [Google Scholar] [CrossRef] [PubMed]
- Inaba, K.; Takahashi, Y.H.; Fujieda, N.; Kano, K.; Miyoshi, H.; Ito, K. DsbB elicits a red-shift of bound ubiquinone during the catalysis of DsbA oxidation. J. Biol. Chem. 2004, 279, 6761–6768. [Google Scholar] [CrossRef] [PubMed]
- Kadokura, H.; Bader, M.; Tian, H.; Bardwell, J.C.; Beckwith, J. Roles of a conserved arginine residue of DsbB in linking protein disulfide-bond-formation pathway to the respiratory chain of Escherichia coli. Proc. Natl. Acad. Sci. USA 2000, 97, 10884–10849. [Google Scholar] [CrossRef] [PubMed]
- Inaba, K.; Takahashi, Y.-h.; Ito, K.; Hayashi, S. Critical role of a thiolate-quinone charge transfer complex and its adduct form in de novo disulfide bond generation by DsbB. Proc. Natl. Acad. Sci. USA 2006, 103, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Messens, J.; Collet, J.-F. Pathways of disulfide bond formation in Escherichia coli. Intl. J. Biochem. Cell Biol. 2006, 38, 1050–1062. [Google Scholar] [CrossRef]
- Hatahet, F.; Boyd, D.; Beckwith, J. Disulfide bond formation in prokaryotes: History, diversity and design. Biochim. Biophys. Acta 2014, 1844, 1402–1414. [Google Scholar] [CrossRef] [PubMed]
- Ito, K.; Inaba, K. The disulfide bond formation (Dsb) system. Curr. Opin. Struct. Biol. 2008, 18, 450–458. [Google Scholar] [CrossRef] [PubMed]
- Daniels, R.; Mellroth, P.; Bernsel, A.; Neiers, F.; Normark, S.; von Heijne, G.; Henriques-Normark, B. Disulfide bond formation and cysteine exclusion in gram-positive bacteria. J. Biol. Chem. 2010, 285, 3300–3309. [Google Scholar] [CrossRef] [PubMed]
- Inaba, K. Disulfide bond formation system in Escherichia coli. J. Biochem. 2009, 146, 591–597. [Google Scholar] [CrossRef] [PubMed]
- Bardwell, J.C.A.; McGovern, K.; Beckwith, J. Identification of a protein required for disulfide bond formation in vivo. Cell 1991, 67, 581–589. [Google Scholar] [CrossRef] [PubMed]
- Grauschopf, U.; Winther, J.R.; Korber, P.; Zander, T.; Dallinger, P.; Bardwell, J.C.A. Why is DsbA such an oxidizing disulfide catalyst? Cell 1995, 83, 947–955. [Google Scholar] [CrossRef] [PubMed]
- Martin, J.L.; Bardwell, J.C.A.; Kuriyan, J. Crystal structure of the DsbA protein required for disulphide bond formation in vivo. Nature 1993, 365, 464–468. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, T.; Kishigami, S.; Sone, M.; Inokuchi, H.; Mogi, T.; Ito, K. Respiratory chain is required to maintain oxidized states of the DsbA-DsbB disulfide bond formation system in aerobically growing Escherichia coli cells. Proc. Natl. Acad. Sci. USA 1997, 94, 11857–11862. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Cierpicki, T.; Jimenez, R.H.; Lukasik, S.M.; Ellena, J.F.; Cafiso, D.S.; Kadokura, H.; Beckwith, J.; Bushweller, J.H. NMR solution structure of the integral membrane enzyme DsbB: Functional insights into DsbB-catalyzed disulfide bond formation. Mol. Cell 2008, 31, 896–908. [Google Scholar] [CrossRef] [PubMed]
- Inaba, K.; Murakami, S.; Suzuki, M.; Nakagawa, A.; Yamashita, E.; Okada, K.; Ito, K. Crystal structure of the DsbB-DsbA complex reveals a mechanism of disulfide bond generation. Cell 2006, 127, 789–801. [Google Scholar] [CrossRef] [PubMed]
- Guddat, L.W.; Bardwell, J.C.; Martin, J.L. Crystal structures of reduced and oxidized DsbA: Investigation of domain motion and thiolate stabilization. Structure 1998, 6, 6757–6767. [Google Scholar] [CrossRef]
- Schirra, H.J.; Renner, C.; Czisch, M.; Huber-Wunderlich, M.; Holak, T.A.; Glockshuber, R. Structure of reduced DsbA from Escherichia coli in solution. Biochemistry 1998, 37, 6263–6276. [Google Scholar] [CrossRef] [PubMed]
- Katzen, F.; Beckwith, J. Transmembrane electron transfer by the membrane protein DsbD occurs via a disulfide bond cascade. Cell 2000, 103, 769–779. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.H.; Porat, A.; Ye, J.; Beckwith, J. Redox-active cysteines of a membrane electron transporter DsbD show dual compartment accessibility. EMBO J. 2007, 26, 3509–3520. [Google Scholar] [CrossRef] [PubMed]
- Rozhkova, A.; Stirnimann, C.U.; Frei, P.; Grauschopf, U.; Brunisholz, R.; Grutter, M.G.; Capitani, G.; Glockshuber, R. Structural basis and kinetics of inter- and intramolecular disulfide exchange in the redox catalyst DsbD. EMBO J. 2004, 23, 1709–1719. [Google Scholar] [CrossRef] [PubMed]
- Nagy, P. Kinetics and mechanisms of thiol-disulfide exchange covering direct substitution and thiol oxidation-mediated pathways. Antioxid. Redox Signal. 2013, 18, 1623–1641. [Google Scholar] [CrossRef] [PubMed]
- Haebel, P.W.; Goldstone, D.; Katzen, F.; Beckwith, J.; Metcalf, P. The disulfide bond isomerase DsbC is activated by an immunoglobulin-fold thiol oxidoreductase: Crystal structure of the DsbC-DsbDalpha complex. EMBO J. 2002, 21, 4774–4784. [Google Scholar] [CrossRef] [PubMed]
- Bulleid, N.J. Disulfide bond formation in the mammalian endoplasmic reticulum. Cold Spring Harb. Perspect. Biol. 2012, 4. [Google Scholar] [CrossRef]
- Arnér, E.S.J. Focus on mammalian thioredoxin reductases—Important selenoproteins with versatile functions. Biochim. Biophys. Acta 2009, 1790, 495–526. [Google Scholar] [CrossRef] [PubMed]
- Hebert, D.N.; Molinari, M. In and out of the ER: Protein folding, quality control, degradation, and related human diseases. Physiol. Rev. 2007, 87, 1377–408. [Google Scholar] [CrossRef] [PubMed]
- Sevier, C.S.; Kaiser, C.A. Conservation and diversity of the cellular disulfide bond formation pathways. Antioxid. Redox Signal. 2006, 8, 797–811. [Google Scholar] [CrossRef] [PubMed]
- Sevier, C.S.; Kaiser, C.A. Formation and transfer of disulphide bonds in living cells. Nat. Rev. Mol. Cell Biol. 2002, 3, 836–847. [Google Scholar] [CrossRef] [PubMed]
- Inaba, K.; Masui, S.; Iida, H.; Vavassori, S.; Sitia, R.; Suzuki, M. Crystal structures of human Ero1alpha reveal the mechanisms of regulated and targeted oxidation of PDI. EMBO J. 2010, 29, 3330–3343. [Google Scholar] [CrossRef] [PubMed]
- Benham, A.M.; Cabibbo, A.; Fassio, A.; Bulleid, N.; Sitia, R.; Braakman, I. The CXXCXXC motif determines the folding, structure and stability of human Ero1-Lalpha. EMBO J. 2000, 19, 4493–4502. [Google Scholar] [CrossRef] [PubMed]
- Edman, J.C.; Ellis, L.; Blacher, R.W.; Roth, R.A.; Rutter, W.J. Sequence of protein disulphide isomerase and implications of its relationship to thioredoxin. Nature 1985, 317, 267–270. [Google Scholar] [CrossRef] [PubMed]
- Gross, E.; Kastner, D.B.; Kaiser, C.A.; Fass, D. Structure of Ero1p, source of disulfide bonds for oxidative protein folding in the cell. Cell 2004, 117, 601–610. [Google Scholar] [CrossRef] [PubMed]
- Appenzeller-Herzog, C.; Riemer, J.; Christensen, B.; Sorensen, E.S.; Ellgaard, L. A novel disulphide switch mechanism in Ero1alpha balances ER oxidation in human cells. EMBO J. 2008, 27, 2977–2987. [Google Scholar] [CrossRef] [PubMed]
- Darby, N.J.; Freedman, R.B.; Creighton, T.E. Dissecting the mechanism of protein disulfide isomerase: Catalysis of disulfide bond formation in a model peptide. Biochemistry 1994, 33, 7937–7947. [Google Scholar] [CrossRef] [PubMed]
- Mesecke, N.; Terziyska, N.; Kozany, C.; Baumann, F.; Neupert, W.; Hell, K.; Herrmann, J.M. A disulfide relay system in the intermembrane space of mitochondria that mediates protein import. Cell 2005, 121, 1059–1069. [Google Scholar] [CrossRef] [PubMed]
- Allen, S.; Balabanidou, V.; Sideris, D.P.; Lisowsky, T.; Tokatlidis, K. Erv1 mediates the Mia40-dependent protein import pathway and provides a functional link to the respiratory chain by shuttling electrons to cytochrome c. J. Mol. Biol. 2005, 353, 937–944. [Google Scholar] [CrossRef] [PubMed]
- Tokatlidis, K. A disulfide relay system in mitochondria. Cell 2005, 121, 965–967. [Google Scholar] [CrossRef] [PubMed]
- Grumbt, B.; Stroobant, V.; Terziyska, N.; Israel, L.; Hell, K. Functional characterization of Mia40p, the central component of the disulfide relay system of the mitochondrial intermembrane space. J. Biol. Chem. 2007, 282, 37461–37470. [Google Scholar] [CrossRef] [PubMed]
- Banci, L.; Bertini, I.; Cefaro, C.; Cenacchi, L.; Ciofi-Baffoni, S.; Felli, I. C.; Gallo, A.; Gonnelli, L.; Luchinat, E.; Sideris, D.; Tokatlidis, K. Molecular chaperone function of Mia40 triggers consecutive induced folding steps of the substrate in mitochondrial protein import. Proc. Natl. Acad. Sci. USA 2010, 107, 20190–20195. [Google Scholar] [CrossRef] [PubMed]
- Milenkovic, D.; Ramming, T.; Müller, J. M.; Wenz, L.-S.; Gebert, N.; Schulze-Specking, A.; Stojanovski, D.; Rospert, S.; Chacinska, A. Identification of the signal directing Tim9 and Tim10 into the intermembrane space of mitochondria. Mol. Biol. Cell 2009, 20, 2530–2539. [Google Scholar] [CrossRef] [PubMed]
- Sideris, D. P.; Petrakis, N.; Katrakili, N.; Mikropoulou, D.; Gallo, A.; Ciofi-Baffoni, S.; Banci, L.; Bertini, I.; Tokatlidis, K. A novel intermembrane space-targeting signal docks cysteines onto Mia40 during mitochondrial oxidative folding. J. Cell. Biol. 2009, 187, 1007–1022. [Google Scholar] [CrossRef] [PubMed]
- Banci, L.; Bertini, I.; Cefaro, C.; Ciofi-Baffoni, S.; Gallo, A.; Martinelli, M.; Sideris, D.P.; Katrakili, N.; Tokatlidis, K. MIA40 is an oxidoreductase that catalyzes oxidative protein folding in mitochondria. Nat. Struct. Mol. Biol. 2009, 16, 198–206. [Google Scholar] [CrossRef] [PubMed]
- Bien, M.; Longen, S.; Wagener, N.; Chwalla, I.; Herrmann, J.M.; Riemer, J. Mitochondrial disulfide bond formation is driven by intersubunit electron transfer in Erv1 and proofread by glutathione. Mol. Cell 2010, 37, 516–528. [Google Scholar] [CrossRef] [PubMed]
- Naoé, M.; Ohwa, Y.; Ishikawa, D.; Ohshima, C.; Nishikawa, S.-I.; Yamamoto, H.; Endo, T. Identification of Tim40 that mediates protein sorting to the mitochondrial intermembrane space. J. Biol. Chem. 2004, 279, 47815–47821. [Google Scholar] [CrossRef] [PubMed]
- Chacinska, A.; Pfannschmidt, S.; Wiedemann, N.; Kozjak, V.; Sanjuan Szklarz, L.K.; Schulze-Specking, A.; Truscott, K.N.; Guiard, B.; Meisinger, C.; Pfanner, N. Essential role of Mia40 in import and assembly of mitochondrial intermembrane space proteins. EMBO J. 2004, 23, 3735–3746. [Google Scholar] [CrossRef] [PubMed]
- Coppock, D.L.; Thorpe, C. Multidomain flavin-dependent sulfhydryl oxidases. Antioxid. Redox Signal. 2006, 8, 300–311. [Google Scholar] [CrossRef] [PubMed]
- Sztolsztener, M.E.; Brewinska, A.; Guiard, B.; Chacinska, A. Disulfide bond formation: Sulfhydryl oxidase ALR controls mitochondrial biogenesis of human MIA40. Traffic 2013, 14, 309–320. [Google Scholar] [CrossRef] [PubMed]
- Faccio, G.; Nivala, O.; Kruus, K.; Buchert, J.; Saloheimo, M. Sulfhydryl oxidases: Sources, properties, production and applications. Appl. Microbiol. Biotechnol. 2011, 91, 957–966. [Google Scholar] [CrossRef] [PubMed]
- Gross, E.; Sevier, C.S.; Vala, A.; Kaiser, C. A.; Fass, D. A new FAD-binding fold and intersubunit disulfide shuttle in the thiol oxidase Erv2p. Nat. Struct. Biol. 2002, 9, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Hatahet, F.; Ruddock, L.W. Protein disulfide isomerase: A critical evaluation of its function in disulfide bond formation. Antioxid. Redox Signal. 2009, 11, 2807–2850. [Google Scholar] [CrossRef] [PubMed]
- Kawagoe, Y.; Suzuki, K.; Tasaki, M.; Yasuda, H.; Akagi, K.; Katoh, E.; Nishizawa, N.K.; Ogawa, M.; Takaiwa, F. The critical role of disulfide bond formation in protein sorting in the endosperm of rice. Plant Cell 2005, 17, 1141–1153. [Google Scholar] [CrossRef] [PubMed]
- Levitan, A.; Danon, A.; Lisowsky, T. Unique features of plant mitochondrial sulfhydryl oxidase. J. Biol. Chem. 2004, 279, 20002–20008. [Google Scholar] [CrossRef] [PubMed]
- Dabir, D.V.; Leverich, E.P.; Kim, S.K.; Tsai, F.D.; Hirasawa, M.; Knaff, D.B.; Koehler, C.M. A role for cytochrome c and cytochrome c peroxidase in electron shuttling from Erv1. EMBO J. 2007, 26, 4801–4811. [Google Scholar] [CrossRef] [PubMed]
- Baena-Gonzalez, E.; Aro, E.M. Biogenesis, assembly and turnover of photosystem II units. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2002, 357, 1451–1459. [Google Scholar] [CrossRef] [PubMed]
- Onda, Y. Oxidative protein-folding systems in plant cells. Int. J. Cell Biol. 2013, 2013. [Google Scholar] [CrossRef]
- Wittenberg, G.; Danon, A. Disulfide bond formation in chloroplasts: Formation of disulfide bonds in signaling chloroplast proteins. Plant Sci. 2008, 175, 459–466. [Google Scholar] [CrossRef]
- Goodstadt, L.; Ponting, C.P. Vitamin K epoxide reductase: homology, active site and catalytic mechanism. Trends Biochem. Sci. 2004, 29, 289–292. [Google Scholar] [CrossRef] [PubMed]
- Tie, J.K.; Jin, D.Y.; Stafford, D.W. Human vitamin K epoxide reductase and its bacterial homologue have different membrane topologies and reaction mechanisms. J. Biol. Chem. 2012, 287, 33945–33955. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Schulman, S.; Dutton, R.J.; Boyd, D.; Beckwith, J.; Rapoport, T.A. Structure of a bacterial homologue of vitamin K epoxide reductase. Nature 2010, 463, 507–512. [Google Scholar] [CrossRef] [PubMed]
- Karamoko, M.; Cline, S.; Redding, K.; Ruiz, N.; Hamel, P.P. Lumen thiol oxidoreductase1, a disulfide bond-forming catalyst, is required for the assembly of photosystem II in Arabidopsis. Plant Cell 2011, 23, 4462–4475. [Google Scholar] [CrossRef] [PubMed]
- Furt, F.; Oostende, C.; Widhalm, J.R.; Dale, M.A.; Wertz, J.; Basset, G.J. A bimodular oxidoreductase mediates the specific reduction of phylloquinone (vitamin K1) in chloroplasts. Plant J. 2010, 64, 38–46. [Google Scholar] [PubMed]
- Schulman, S.; Wang, B.; Li, W.; Rapoport, T.A. Vitamin K epoxide reductase prefers ER membrane-anchored thioredoxin-like redox partners. Proc. Natl. Acad. Sci. USA 2010, 107, 15027–15032. [Google Scholar] [CrossRef] [PubMed]
- Lu, Y.; Hall, D.A.; Last, R.L. A small zinc finger thylakoid protein plays a role in maintenance of photosystem II in Arabidopsis thaliana. Plant Cell 2011, 1861–1875. [Google Scholar]
- Muranaka, A.; Watanabe, S.; Sakamoto, A.; Shimada, H. Arabidopsis cotyledon chloroplast biogenesis factor CYO1 uses glutathione as an electron donor and interacts with PSI (A1 and A2) and PSII (CP43 and CP47) subunits. J. Plant Physiol. 2012, 169, 1212–1215. [Google Scholar] [CrossRef] [PubMed]
- Argyrou, A.; Blanchard, J.S. Flavoprotein disulfide reductases: Advances in chemistry and function. Prog. Nucleic Acid Res. Mol. Biol. 2004, 78, 89–142. [Google Scholar] [PubMed]
- Chivers, P.T.; Laboissiere, M.C.; Raines, R.T. The CXXC motif: imperatives for the formation of native disulfide bonds in the cell. EMBO J. 1996, 15, 2659–2667. [Google Scholar] [PubMed]
- Heckler, E.J.; Rancy, P.C.; Kodali, V.K.; Thorpe, C. Generating disulfides with the Quiescin-sulfhydryl oxidases. Biochim. Biophys. Acta 2008, 1783, 567–577. [Google Scholar] [CrossRef] [PubMed]
- Thorpe, C.; Hoober, K.L.; Raje, S.; Glynn, N.M.; Burnside, J.; Turi, G.K.; Coppock, D.L. Sulfhydryl oxidases: emerging catalysts of protein disulfide bond formation in eukaryotes. Arch. Biochem. Biophys. 2002, 405, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Raje, S.; Thorpe, C. Inter-domain redox communication in flavoenzymes of the quiescin/sulfhydryl oxidase family: Role of a thioredoxin domain in disulfide bond formation. Biochemistry 2003, 42, 4560–4568. [Google Scholar] [CrossRef] [PubMed]
- Meyer, S.; Lorenz, C.; Baser, B.; Wordehoff, M.; Jager, V.; van den Heuvel, J. Multi-host expression system for recombinant production of challenging proteins. PLoS One 2013, 8, e68674. [Google Scholar] [CrossRef] [PubMed]
- Overton, T.W. Recombinant protein production in bacterial hosts. Drug Discov. Today 2014, 19, 590–601. [Google Scholar] [CrossRef] [PubMed]
- Laitinen, O.H.; Airenne, K.J.; Hytonen, V.P.; Peltomaa, E.; Mahonen, A.J.; Wirth, T.; Lind, M.M.; Makela, K.A.; Toivanen, P.I.; Schenkwein, D.; Heikura, T.; Nordlund, H.R.; Kulomaa, M.S.; Yla-Herttuala, S. A multipurpose vector system for the screening of libraries in bacteria, insect and mammalian cells and expression in vivo. Nucleic Acids Res. 2005, 33, e42. [Google Scholar] [CrossRef] [PubMed]
- De Marco, A. Recombinant polypeptide production in E. coli: Towards a rational approach to improve the yields of functional proteins. Microb. Cell Fact. 2013, 12. [Google Scholar] [CrossRef]
- Liu, L.; Yang, H.; Shin, H.D.; Chen, R.R.; Li, J.; Du, G.; Chen, J. How to achieve high-level expression of microbial enzymes: Strategies and perspectives. Bioengineered 2013, 4, 212–223. [Google Scholar] [CrossRef] [PubMed]
- Brondyk, W.H. Selecting an appropriate method for expressing a recombinanat protein. In Guide to Protein Purification, 2nd ed.; Burgess, R.R., Deutscher, M.P., Eds.; Gulf Professional Publishing: San Diego, CA, USA, 2009; Volume 463, p. 131. [Google Scholar]
- Klint, J.K.; Senff, S.; Saez, N.J.; Seshadri, R.; Lau, H.Y.; Bende, N.S.; Undheim, E.A.; Rash, L.D.; Mobli, M.; King, G.F. Production of recombinant disulfide-rich venom peptides for structural and functional analysis via expression in the periplasm of E. coli. PLoS One 2013, 8, e63865. [Google Scholar] [CrossRef] [PubMed]
- Nakamoto, H.; Bardwell, J.C. Catalysis of disulfide bond formation and isomerization in the Escherichia coli periplasm. Biochim. Biophys. Acta 2004, 1694, 111–119. [Google Scholar] [CrossRef] [PubMed]
- Gorman, C.M.; Groskreutz, D.J.; Marriott, D. Method and host cells for producing two-chain relaxin polypeptide. Eur. Patent 1,099,758, 16 May 2001. [Google Scholar]
- Chen, X.; Bai, Y.; Zaro, J.L.; Shen, W.C. Design of an in vivo cleavable disulfide linker in recombinant fusion proteins. BioTechniques 2010, 49, 513–518. [Google Scholar] [CrossRef] [PubMed]
- Jia, X.Y.; Guo, Z.Y.; Wang, Y.; Xu, Y.; Duan, S.S.; Feng, Y.M. Peptide models of four possible insulin folding intermediates with two disulfides. Protein Sci. 2003, 12, 2412–2419. [Google Scholar] [CrossRef] [PubMed]
- Bessette, P.H.; Aslund, F.; Beckwith, J.; Georgiou, G. Efficient folding of proteins with multiple disulfide bonds in the Escherichia coli cytoplasm. Proc. Natl. Acad. Sci. USA 1999, 96, 13703–13708. [Google Scholar] [CrossRef] [PubMed]
- Berkmen, M. Production of disulfide-bonded proteins in Escherichia coli. Protein Expr. Purif. 2012, 82, 240–251. [Google Scholar] [CrossRef] [PubMed]
- Walsh, G. Biopharmaceutical benchmarks 2010. Nat. Biotechnol. 2010, 28, 917–924. [Google Scholar] [CrossRef] [PubMed]
- Walsh, G. Biopharmaceutical benchmarks 2014. Nat. Biotechnol. 2014, 32, 992–1000. [Google Scholar] [CrossRef] [PubMed]
- Clark, E.D.B. Refolding of recombinant proteins. Curr. Opin. Biotechnol. 1998, 9, 9157–9163. [Google Scholar]
- Rudolph, R.; Lilie, H. In vitro folding of inclusion body proteins. FASEB J. 1996, 10, 49–56. [Google Scholar] [PubMed]
- Helenbeck, R.; Koths, K.E.; Cowgill, C.; Laird, W.J. Production of purified, biologically active, bacterially produced recombinant human CSF-1. US Patent 4,929,700, 29 May 1990. [Google Scholar]
- Koths, K.E.; Halenbeck, R.F. Method for promoting disulfide bond formation in recombinant proteins. US Patent 4,572,798, 25 February 1986. [Google Scholar]
- Freedman, R.B. The formation of protein disulphide bonds. Curr. Opin. Struct. Biol. 1995, 5, 85–91. [Google Scholar] [CrossRef] [PubMed]
- Frand, A.R.; Cuozzo, J.W.; Kaiser, C.A. Pathways for protein disulphide bond formation. Trends Cell Biol. 2000, 10, 203–210. [Google Scholar] [CrossRef] [PubMed]
- Luan, C.; Zhang, H.; Song, D.; Xie, Y.; Feng, J.; Wang, Y. Expressing antimicrobial peptide cathelicidin-BF in Bacillus subtilis using SUMO technology. Appl. Microbiol. Biotechnol. 2014, 98, 3651–3658. [Google Scholar] [CrossRef] [PubMed]
- Alibolandi, M.; Mirzahoseini, H. Chemical assistance in refolding of bacterial inclusion bodies. Biochem. Res. Int. 2011, 2011. [Google Scholar] [CrossRef]
- Sandee, D.; Tungpradabkul, S.; Kurokawa, Y.; Fukui, K.; Takagi, M. Combination of Dsb coexpression and an addition of sorbitol markedly enhanced soluble expression of single-chain Fv in Escherichia coli. Biotechnol. Bioeng. 2005, 91, 418–424. [Google Scholar] [CrossRef] [PubMed]
- Kondo, A.; Kohda, J.; Endo, Y.; Shiromizu, T.; Kurokawa, Y.; Nishihara, K.; Yanagi, H.; Yura, T.; Fukuda, H. Improvement of productivity of active horseradish peroxidase in Escherichia coli by coexpression of Dsb proteins. J. Biosci. Bioeng. 2000, 90, 600–606. [Google Scholar] [CrossRef] [PubMed]
- Derman, A.; Prinz, W.; Belin, D.; Beckwith, J. Mutations that allow disulfide bond formation in the cytoplasm of Escherichia coli. Science 1993, 262, 1744–1747. [Google Scholar] [CrossRef] [PubMed]
- Kaomek, M.; Mizuno, K.; Fujimura, T.; Sriyotha, P.; Cairns, J.R.K. Cloning, expression, and characterization of an antifungal chitinase from Leucaena leucocephala de Wit. Biosci. Biotechnol. Biochem. 2003, 67, 667–676. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, T.; Shibata, H.; Araya, T.; Nakatsu, T.; Miyairi, K.; Okuno, T.; Kato, H. Expression, purification, and crystallization of endopolygalacturonase from a pathogenic fungus, Stereum purpureum, in Escherichia coli. Protein Expr. Purif. 2005, 44, 130–135. [Google Scholar] [CrossRef] [PubMed]
- Bar, M.; Bar-Ziv, R.; Scherf, T.; Fass, D. Efficient production of a folded and functional, highly disulfide-bonded β-helix antifreeze protein in bacteria. Protein Expr. Purif. 2006, 48, 243–252. [Google Scholar] [CrossRef] [PubMed]
- Berndt, C.; Lillig, C.H.; Holmgren, A. Thioredoxins and glutaredoxins as facilitators of protein folding. Biochim. Biophys. Acta 2008, 1783, 641–650. [Google Scholar] [CrossRef] [PubMed]
- Jurado, P.; de Lorenzo, V.; Fernandez, L.A. Thioredoxin fusions increase folding of single chain Fv antibodies in the cytoplasm of Escherichia coli: Evidence that chaperone activity is the prime effect of thioredoxin. J. Mol. Biol. 2006, 357, 49–61. [Google Scholar] [CrossRef] [PubMed]
- Heo, M.A.; Kim, S.H.; Kim, S.Y.; Kim, Y.J.; Chung, J.; Oh, M.K.; Lee, S.G. Functional expression of single-chain variable fragment antibody against c-Met in the cytoplasm of Escherichia coli. Protein Expr. Purif. 2006, 47, 203–209. [Google Scholar] [CrossRef] [PubMed]
- De Marco, A. Strategies for successful recombinant expression of disulfide bond-dependent proteins in Escherichia coli. Microb. Cell Fact. 2009, 8. [Google Scholar] [CrossRef] [PubMed]
- Huber, D.; Boyd, D.; Xia, Y.; Olma, M. H.; Gerstein, M.; Beckwith, J. Use of thioredoxin as a reporter to identify a subset of Escherichia coli signal sequences that promote signal recognition particle-dependent translocation. J. Bacteriol. 2005, 187, 2983–2991. [Google Scholar] [CrossRef] [PubMed]
- Mergulhao, F.J.; Monteiro, G.A.; Larsson, G.; Bostrom, M.; Farewell, A.; Nystrom, T.; Cabral, J.M.; Taipa, M.A. Evaluation of inducible promoters on the secretion of a ZZ-proinsulin fusion protein in Escherichia coli. Biotechnol. Appl. Biochem. 2003, 38, 87–93. [Google Scholar] [CrossRef] [PubMed]
- Obukowicz, M.G.; Gustafson, M.E.; Junger, K.D.; Leimgruber, R.M.; Wittwer, A.J.; Wun, T.C.; Warren, T.G.; Bishop, B.F.; Mathis, K.J.; McPherson, D.T.; et al. Secretion of active kringle-2-serine protease in Escherichia coli. Biochemistry 1990, 29, 9737–9745. [Google Scholar] [CrossRef] [PubMed]
- Pasek, M.; Boeggeman, E.; Ramakrishnan, B.; Qasba, P.K. Galectin-1 as a fusion partner for the production of soluble and folded human beta-1,4-galactosyltransferase-T7 in E. coli. Biochem. Biophys. Res. Commun. 2010, 394, 679–684. [Google Scholar] [CrossRef] [PubMed]
- Thie, H.; Schirrmann, T.; Paschke, M.; Dübel, S.; Hust, M. SRP and Sec pathway leader peptides for antibody phage display and antibody fragment production in E. coli. New Biotechnol. 2008, 25, 49–54. [Google Scholar] [CrossRef]
- Malik, A.; Rudolph, R.; Söhling, B. A novel fusion protein system for the production of native human pepsinogen in the bacterial periplasm. Protein Expr. Purif. 2006, 47, 662–671. [Google Scholar] [CrossRef] [PubMed]
- Maskos, K.; Huber-Wunderlich, M.; Glockshuber, R. DsbA and DsbC-catalyzed oxidative folding of proteins with complex disulfide bridge patterns in vitro and in vivo. J. Mol. Biol. 2003, 325, 495–513. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Yasin, A.; Tang, R.; Scharer, J.; Moo-Young, M.; Chou, C.P. Heterologous expression of lipase in Escherichia coli is limited by folding and disulfide bond formation. Appl. Microbiol. Biotechnol. 2008, 81, 79–87. [Google Scholar] [CrossRef] [PubMed]
- Sklar, J.G. Defining the roles of the periplasmic chaperones SurA, Skp, and DegP in Escherichia coli. Genes Dev. 2007, 21, 2473–2484. [Google Scholar] [CrossRef] [PubMed]
- Knoblauch, N.T.; Rudiger, S.; Schonfeld, H.J.; Driessen, A.J.; Schneider-Mergener, J.; Bukau, B. Substrate specificity of the SecB chaperone. J. Biol. Chem. 1999, 274, 34219–34225. [Google Scholar] [CrossRef] [PubMed]
- Mapa, K.; Tiwari, S.; Kumar, V.; Jayaraj, G.G.; Maiti, S. Information encoded in non-native states drives substrate-chaperone pairing. Structure 2012, 20, 1562–1573. [Google Scholar] [CrossRef] [PubMed]
- Martin, C.; Rojas, G.; Mitchell, J.; Vincent, K.; Wu, J.; McCafferty, J.; Schofield, D. A simple vector system to improve performance and utilisation of recombinant antibodies. BMC Biotechnol. 2006, 6. [Google Scholar] [CrossRef]
- Schmoldt, H.-U.; Wentzel, A.; Becker, S.; Kolmar, H. A fusion protein system for the recombinant production of short disulfide bond rich cystine knot peptides using barnase as a purification handle. Protein Expr. Purif. 2005, 39, 82–89. [Google Scholar] [CrossRef] [PubMed]
- Niemann, H.H.; Schmoldt, H.-U.; Wentzel, A.; Kolmar, H.; Heinz, D.W. Barnase fusion as a tool to determine the crystal structure of the small disulfide-rich protein McoEeTI. J. Mol. Biol. 2006, 356, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Schäffner, J.; Winter, J.; Rudolph, R.; Schwarz, E. Cosecretion of chaperones and low-molecular-size medium additives increases the yield of recombinant disulfide-bridged proteins. Appl. Environ. Microbiol. 2001, 67, 3994–4000. [Google Scholar] [CrossRef] [PubMed]
- Mogk, A.; Mayer, M.P.; Deuerling, E. Mechanisms of protein folding: molecular chaperones and their application in biotechnology. ChemBioChem 2002, 3, 807–814. [Google Scholar] [CrossRef] [PubMed]
- Bandyopadhyay, A.; Saxena, K.; Kasturia, N.; Dalal, V.; Bhatt, N.; Rajkumar, A.; Maity, S.; Sengupta, S.; Chakraborty, K. Chemical chaperones assist intracellular folding to buffer mutational variations. Nat. Chem. Biol. 2012, 8, 238–245. [Google Scholar] [CrossRef] [PubMed]
- Barth, S.; Huhn, M.; Matthey, B.; Klimka, A.; Galinski, E.A.; Engert, A. Compatible-solute-supported periplasmic expression of functional recombinant proteins under stress conditions. Appl. Environ. Microbiol. 2000, 66, 1572–1579. [Google Scholar] [CrossRef] [PubMed]
- Andrews, H.; Li, Z.; Altuve-Blanco, A.; Rivera, M.; Burnap, R.L. Expression, mutagenesis, and characterization of recombinant low-potential cytochrome c550 of photosystem II. Biochemistry 2005, 44, 6092–6100. [Google Scholar] [CrossRef] [PubMed]
- Winter, J.; Neubauer, P.; Glockshuber, R.; Rudolph, R. Increased production of human proinsulin in the periplasmic space of Escherichia coli by fusion to DsbA. J. Biotechnol. 2001, 84, 175–185. [Google Scholar] [CrossRef] [PubMed]
- Shimamoto, S.; Katayama, H.; Okumura, M.; Hidaka, Y. Chemical methods and approaches to the regioselective formation of multiple disulfide bonds. Curr. Protoc. Protein Sci. 2014, 76, 28.8.1–28.8.28. [Google Scholar] [PubMed]
- Okumura, M.; Shimamoto, S.; Hidaka, Y. Chemical methods for producing disulfide bonds in peptides and proteins to study folding regulation. Curr. Protoc. Protein Sci. 2014, 76, 28.7.1–28.7.13. [Google Scholar] [PubMed]
- Hidaka, Y. Overview of the regulation of disulfide bond formation in Peptide and protein folding. Curr. Protoc. Protein Sci. 2014, 76, 28.6.1–28.6.6. [Google Scholar] [PubMed]
- Harada, T.; Kurimoto, E.; Moriyama, Y.; Ejima, D.; Sakai, T.; Nohara, D.; Kato, K. Application of combined reagent solution to the oxidative refolding of recombinant human interleukin 6. Chem. Pharm. Bull. 2001, 49, 1128–1131. [Google Scholar] [CrossRef] [PubMed]
- Ito, L.; Okumura, M.; Tao, K.; Kasai, Y.; Tomita, S.; Oosuka, A.; Yamada, H.; Shibano, T.; Shiraki, K.; Kumasaka, T.; et al. Glutathione ethylester, a novel protein refolding reagent, enhances both the efficiency of refolding and correct disulfide formation. Protein J. 2012, 31, 499–503. [Google Scholar] [CrossRef] [PubMed]
- Arai, K.; Noguchi, M.; Singh, B.G.; Priyadarsini, K.I.; Fujio, K.; Kubo, Y.; Takayama, K.; Ando, S.; Iwaoka, M. A water-soluble selenoxide reagent as a useful probe for the reactivity and folding of polythiol peptides. FEBS Open Biol. 2013, 3, 55–64. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Patil, N.A.; Tailhades, J.; Hughes, R.A.; Separovic, F.; Wade, J.D.; Hossain, M.A. Cellular Disulfide Bond Formation in Bioactive Peptides and Proteins. Int. J. Mol. Sci. 2015, 16, 1791-1805. https://doi.org/10.3390/ijms16011791
Patil NA, Tailhades J, Hughes RA, Separovic F, Wade JD, Hossain MA. Cellular Disulfide Bond Formation in Bioactive Peptides and Proteins. International Journal of Molecular Sciences. 2015; 16(1):1791-1805. https://doi.org/10.3390/ijms16011791
Chicago/Turabian StylePatil, Nitin A., Julien Tailhades, Richard Anthony Hughes, Frances Separovic, John D. Wade, and Mohammed Akhter Hossain. 2015. "Cellular Disulfide Bond Formation in Bioactive Peptides and Proteins" International Journal of Molecular Sciences 16, no. 1: 1791-1805. https://doi.org/10.3390/ijms16011791
APA StylePatil, N. A., Tailhades, J., Hughes, R. A., Separovic, F., Wade, J. D., & Hossain, M. A. (2015). Cellular Disulfide Bond Formation in Bioactive Peptides and Proteins. International Journal of Molecular Sciences, 16(1), 1791-1805. https://doi.org/10.3390/ijms16011791