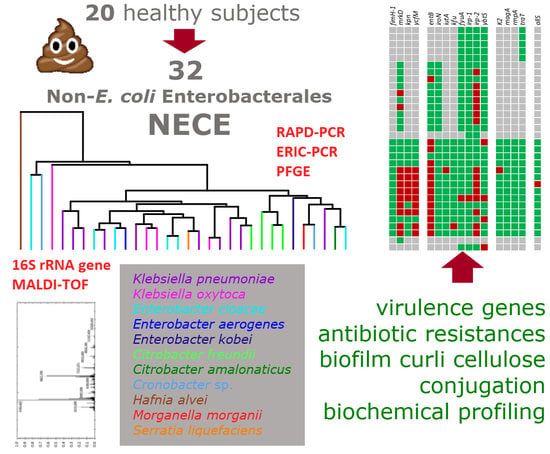
Antibiotic Resistance, Virulence Factors, Phenotyping, and Genotyping of Non-Escherichia coli Enterobacterales from the Gut Microbiota of Healthy Subjects
Abstract
:1. Introduction
2. Results
2.1. Counting and Isolation of NECE
2.2. Taxonomic Attribution and PFGE Genotyping
2.3. Virulence Genotyping
2.4. Biofilm Formation and Production of Curli and Cellulose
2.5. Conjugation
2.6. Antibiotic Resistance
2.7. Biochemical Characterization
3. Discussion
4. Materials and Methods
4.1. Isolation and Enumeration of Enterobacterales
4.2. PFGE Genotyping
4.3. Taxonomic Attribution
4.4. Profiling of Virulence Genes
4.5. Biofilm and Phenotype Assays
4.6. Solid Mating Conjugation Experiments
4.7. Antibiotic Susceptibility
4.8. Biochemical Characterization
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| NECE | non-E. coli Enterobacterales |
| HCCA | HiCrome Coliform Agar |
| MALDI-TOF MS | Matrix-assisted laser desorption ionization–time of flight mass spectrometry |
| ERIC | Enterobacterial repetitive intergenic consensus |
| RAPD | Random Amplification of Polymorphic DNA |
| UPGMA | Unweighted pair group method with arithmetic means |
References
- Leimbach, A.; Hacker, J.; Dobrindt, U. E. coli as an all-rounder: The thin line between commensalism and pathogenicity. Curr. Top. Microbiol. Immunol. 2013, 358, 3–32. [Google Scholar] [CrossRef] [PubMed]
- Manges, A.R.; Geum, H.M.; Guo, A.; Edens, T.J.; Fibke, C.D.; Pitout, J.D.D. Global extraintestinal pathogenic Escherichia coli (ExPEC) lineages. Clin. Microbiol. Rev. 2019, 32. [Google Scholar] [CrossRef] [PubMed]
- Clermont, O.; Christenson, J.K.; Denamur, E.; Gordon, D.M. The Clermont Escherichia coli phylo-typing method revisited: Improvement of specificity and detection of new phylo-groups. Environ. Microbiol. Rep. 2013, 5, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Croxen, M.A.; Law, R.J.; Scholz, R.; Keeney, K.M.; Wlodarska, M.; Finlay, B.B. Recent advances in understanding enteric pathogenic Escherichia coli. Clin. Microbiol. Rev. 2013, 26, 822–880. [Google Scholar] [CrossRef] [Green Version]
- Raimondi, S.; Righini, L.; Candeliere, F.; Musmeci, E.; Bonvicini, F.; Gentilomi, G.; Starčič Erjavec, M.; Amaretti, A.; Rossi, M. Antibiotic resistance, virulence factors, phenotyping, and genotyping of E. coli isolated from the feces of healthy subjects. Microorganisms 2019, 7, 251. [Google Scholar] [CrossRef] [Green Version]
- Sarowska, J.; Futoma-Koloch, B.; Jama-Kmiecik, A.; Frej-Madrzak, M.; Ksiazczyk, M.; Bugla-Ploskonska, G.; Choroszy-Krol, I. Virulence factors, prevalence and potential transmission of extraintestinal pathogenic Escherichia coli isolated from different sources: Recent reports. Gut Pathog. 2019, 11, 10. [Google Scholar] [CrossRef] [Green Version]
- Martin, R.M.; Bachman, M.A. Colonization, Infection, and the accessory genome of Klebsiella pneumoniae. Front. Cell. Infect. Microbiol. 2018, 8, 4. [Google Scholar] [CrossRef] [Green Version]
- Davin-Regli, A.; Pagès, J.M. Enterobacter aerogenes and Enterobacter cloacae; versatile bacterial pathogens confronting antibiotic treatment. Front. Microbiol. 2015, 6, 392. [Google Scholar] [CrossRef] [Green Version]
- Liu, L.; Chen, D.; Liu, L.; Lan, R.; Hao, S.; Jin, W.; Sun, H.; Wang, Y.; Liang, Y.; Xu, J. Genetic diversity, multidrug resistance, and virulence of Citrobacter freundii from diarrheal patients and healthy individuals. Front. Cell. Infect. Microbiol. 2018, 8, 233. [Google Scholar] [CrossRef] [Green Version]
- Gajdács, M.; Urbán, E. Resistance Trends and epidemiology of citrobacter-enterobacter-serratia in urinary tract infections of inpatients and outpatients (RECESUTI): A 10-year survey. Medicina (Kaunas) 2019, 55, 285. [Google Scholar] [CrossRef] [Green Version]
- Lee, C.R.; Lee, J.H.; Park, K.S.; Jeon, J.H.; Kim, Y.B.; Cha, C.J.; Jeong, B.C.; Lee, S.H. Antimicrobial resistance of hypervirulent Klebsiella pneumoniae: Epidemiology, hypervirulence-associated determinants, and resistance mechanisms. Front. Cell. Infect. Microbiol. 2017, 7, 483. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Azevedo, P.A.A.; Furlan, J.P.R.; Oliveira-Silva, M.; Nakamura-Silva, R.; Gomes, C.N.; Costa, K.R.C.; Stehling, E.G.; Pitondo-Silva, A. Detection of virulence and β-lactamase encoding genes in Enterobacter aerogenes and Enterobacter cloacae clinical isolates from Brazil. Braz. J. Microbiol. 2018, 49 (Suppl. 1), 224–228. [Google Scholar] [CrossRef] [PubMed]
- Mokracka, J.; Koczura, R.; Kaznowski, A. Yersiniabactin and other siderophores produced by clinical isolates of Enterobacter spp. and Citrobacter spp. FEMS Immunol. Med. Microbiol. 2004, 40, 51–55. [Google Scholar] [CrossRef] [Green Version]
- Yuan, C.; Yin, Z.; Wang, J.; Qian, C.; Wei, Y.; Zhang, S.; Jiang, L.; Liu, B. Comparative genomic analysis of Citrobacter and key genes essential for the pathogenicity of Citrobacter koseri. Front. Microbiol. 2019, 10, 2774. [Google Scholar] [CrossRef]
- Cruz, A.; Xicohtencatl-Cortes, J.; González-Pedrajo, B.; Bobadilla, M.; Eslava, C.; Rosas, I. Virulence traits in Cronobacter species isolated from different sources. Can. J. Microbiol. 2011, 57, 735–744. [Google Scholar] [CrossRef]
- Zogaj, X.; Bokranz, W.; Nimtz, M.; Römling, U. Production of cellulose and curli fimbriae by members of the family Enterobacteriaceae isolated from the human gastrointestinal tract. Infect. Immun. 2003, 71, 4151–4158. [Google Scholar] [CrossRef] [Green Version]
- Tursi, S.A.; Tükel, Ç. Curli-containing enteric biofilms inside and out: Matrix composition, immune recognition, and disease implications. Microbiol. Mol. Biol. Rev. 2018, 82. [Google Scholar] [CrossRef] [Green Version]
- Partridge, S.R. Resistance mechanisms in Enterobacteriaceae. Pathology 2015, 47, 276–284. [Google Scholar] [CrossRef]
- El Fertas-Aissani, R.; Messai, Y.; Alouache, S.; Bakour, R. Virulence profiles and antibiotic susceptibility patterns of Klebsiella pneumoniae strains isolated from different clinical specimens. Pathol. Biol. (Paris) 2013, 61, 209–216. [Google Scholar] [CrossRef]
- Compain, F.; Babosan, A.; Brisse, S.; Genel, N.; Audo, J.; Ailloud, F.; Kassis-Chikhani, N.; Arlet, G.; Decré, D. Multiplex PCR for detection of seven virulence factors and K1/K2 capsular serotypes of Klebsiella pneumoniae. J. Clin. Microbiol. 2014, 52, 4377–4380. [Google Scholar] [CrossRef] [Green Version]
- Johnson, J.R.; Porter, S.; Johnston, B.; Kuskowski, M.A.; Spurbeck, R.R.; Mobley, H.L.; Williamson, D.A. Host characteristics and bacterial traits predict experimental virulence for Escherichia coli bloodstream isolates from patients with urosepsis. Open Forum Infect. Dis. 2015, 2. [Google Scholar] [CrossRef] [PubMed]
- Davis, T.J.; Matsen, J.M. Prevalence and characteristics of Klebsiella species: Relation to association with a hospital environment. J. Infect. Dis. 1974, 130, 402–405. [Google Scholar] [CrossRef] [PubMed]
- Lau, H.Y.; Huffnagle, G.B.; Moore, T.A. Host and microbiota factors that control Klebsiella pneumoniae mucosal colonization in mice. Microbes Infect. 2008, 10, 1283–1290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Magill, S.S.; Edwards, J.R.; Bamberg, W.; Beldavs, Z.G.; Dumyati, G.; Kainer, M.A.; Lynfield, R.; Maloney, M.; McAllister-Hollod, L.; Nadle, J.; et al. Multistate point-prevalence survey of health care-associated infections. N. Engl. J. Med. 2014, 370, 1198–1208. [Google Scholar] [CrossRef] [Green Version]
- Lan, Y.; Zhou, M.; Jian, Z.; Yan, Q.; Wang, S.; Liu, W. Prevalence of pks gene cluster and characteristics of Klebsiella pneumoniae-induced bloodstream infections. J. Clin. Lab. Anal. 2019, 33, e22838. [Google Scholar] [CrossRef] [Green Version]
- Lawlor, M.S.; O’connor, C.; Miller, V.L. Yersiniabactin is a virulence factor for Klebsiella pneumoniae during pulmonary infection. Infect. Immun. 2007, 75, 1463–1472. [Google Scholar] [CrossRef] [Green Version]
- Schubert, S.; Cuenca, S.; Fischer, D.; Heesemann, J. High-pathogenicity island of Yersinia pestis in Enterobacteriaceae isolated from blood cultures and urine samples: Prevalence and functional expression. J. Infect. Dis. 2000, 182, 1268–1271. [Google Scholar] [CrossRef] [Green Version]
- Schubert, S.; Rakin, A.; Heesemann, J. The Yersinia high-pathogenicity island (HPI): Evolutionary and functional aspects. Int. J. Med. Microbiol. 2004, 294, 83–94. [Google Scholar] [CrossRef]
- Raffatellu, M. Learning from bacterial competition in the host to develop antimicrobials. Nat. Med. 2018, 24, 1097–1103. [Google Scholar] [CrossRef]
- Fang, C.T.; Chuang, Y.P.; Shun, C.T.; Chang, S.C.; Wang, J.T. A novel virulence gene in Klebsiella pneumoniae strains causing primary liver abscess and septic metastatic complications. J. Exp. Med. 2004, 199, 697–705. [Google Scholar] [CrossRef] [Green Version]
- Yu, W.L.; Ko, W.C.; Cheng, K.C.; Lee, H.C.; Ke, D.S.; Lee, C.C.; Fung, C.P.; Chuang, Y.C. Association between rmpA and magA genes and clinical syndromes caused by Klebsiella pneumoniae in Taiwan. Clin. Infect. Dis. 2006, 42, 1351–1358. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johansson, M.E.; Sjovall, H.; Hansson, G.C. The gastrointestinal mucus system in health and disease. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 352–361. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Flores-Mireles, A.L.; Walker, J.N.; Caparon, M.; Hultgren, S.J. Urinary tract infections: Epidemiology, mechanisms of infection and treatment options. Nat. Rev. Microbiol. 2015, 13, 269–284. [Google Scholar] [CrossRef] [PubMed]
- Hobley, L.; Harkins, C.; MacPhee, C.E.; Stanley-Wall, N.R. Giving structure to the biofilm matrix: An overview of individual strategies and emerging common themes. FEMS Microbiol. Rev. 2015, 39, 649–669. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martin, R.M.; Cao, J.; Brisse, S.; Passet, V.; Wu, W.; Zhao, L.; Malani, P.N.; Rao, K.; Bachman, M.A. Molecular epidemiology of colonizing and infecting isolates of Klebsiella pneumoniae. mSphere 2016, 1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gorrie, C.L.; Mirceta, M.; Wick, R.R.; Edwards, D.J.; Thomson, N.R.; Strugnell, R.A.; Pratt, N.F.; Garlick, J.S.; Watson, K.M.; Pilcher, D.V.; et al. Gastrointestinal carriage is a major reservoir of Klebsiella pneumoniae infection in intensive care patients. Clin. Infect. Dis. 2017, 65, 208–215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gajdács, M.; Bátori, Z.; Ábrók, M.; Lázár, A.; Burián, K. Characterization of resistance in gram-negative urinary isolates using existing and novel indicators of clinical relevance: A 10-year data analysis. Life (Basel) 2020, 10, 16. [Google Scholar] [CrossRef] [Green Version]
- Carrie, C.; Walewski, V.; Levy, C.; Alexandre, C.; Baleine, J.; Charreton, C.; Coche-Monier, B.; Caeymaex, L.; Lageix, F.; Lorrot, M.; et al. Klebsiella pneumoniae and Klebsiella oxytoca meningitis in infants. Epidemiological and clinical features. Arch. Pediatr. 2019, 26, 12–15. [Google Scholar] [CrossRef]
- Pötgens, S.A.; Brossel, H.; Sboarina, M.; Catry, E.; Cani, P.D.; Neyrinck, A.M.; Delzenne, N.M.; Bindels, L.B. Klebsiella oxytoca expands in cancer cachexia and acts as a gut pathobiont contributing to intestinal dysfunction. Sci. Rep. 2018, 8, 12321. [Google Scholar] [CrossRef] [Green Version]
- Gajdács, M.; Ábrók, M.; Lázár, A.; Burián, K. Comparative epidemiology and resistance trends of common urinary pathogens in a tertiary-care hospital: A 10-year surveillance study. Medicina (Kaunas) 2019, 55, 356. [Google Scholar] [CrossRef] [Green Version]
- MacPherson, C.W.; Mathieu, O.; Tremblay, J.; Champagne, J.; Nantel, A.; Girard, S.A.; Tompkins, T.A. Gut bacterial microbiota and its resistome rapidly recover to basal state levels after short-term amoxicillin-clavulanic acid treatment in healthy adults. Sci. Rep. 2018, 8, 11192. [Google Scholar] [CrossRef] [PubMed]
- Leclercq, R.; Cantón, R.; Brown, D.F.; Giske, C.G.; Heisig, P.; MacGowan, A.P.; Mouton, J.W.; Nordmann, P.; Rodloff, A.C.; Rossolini, G.M.; et al. EUCAST expert rules in antimicrobial susceptibility testing. Clin. Microbiol. Infect. 2013, 19, 141–160. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wilson, L.A.; Sharp, P.M. Enterobacterial repetitive intergenic consensus (ERIC) sequences in Escherichia coli: Evolution and implications for ERIC-PCR. Mol. Biol. Evol. 2006, 23, 1156–1168. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Quartieri, A.; Simone, M.; Gozzoli, C.; Popovic, M.; D’Auria, G.; Amaretti, A.; Raimondi, S.; Rossi, M. Comparison of culture-dependent and independent approaches to characterize fecal bifidobacteria and lactobacilli. Anaerobe 2016, 38, 130–137. [Google Scholar] [CrossRef]
- Raimondi, S.; Luciani, R.; Sirangelo, T.M.; Amaretti, A.; Leonardi, A.; Ulrici, A.; Foca, G.; D’Auria, G.; Moya, A.; Zuliani, V.; et al. Microbiota of sliced cooked ham packaged in modified atmosphere throughout the shelf life: Microbiota of sliced cooked ham in MAP. Int. J. Food Microbiol. 2019, 16, 200–208. [Google Scholar] [CrossRef]
- Mencacci, A.; Monari, C.; Leli, C.; Merlini, L.; De Carolis, E.; Vella, A.; Cacioni, M.; Buzi, S.; Nardelli, E.; Bistoni, F.; et al. Typing of nosocomial outbreaks of Acinetobacter baumannii by use of matrix-assisted laser desorption ionization-time of flight mass spectrometry. J. Clin. Microbiol. 2013, 51, 603–606. [Google Scholar] [CrossRef] [Green Version]
- Sicard, J.F.; Vogeleer, P.; Le Bihan, G.; Rodriguez Olivera, Y.; Beaudry, F.; Jacques, M.; Harel, J. N-Acetyl-glucosamine influences the biofilm formation of Escherichia coli. Gut Pathog. 2018, 22, 10–26. [Google Scholar] [CrossRef]
- Vogeleer, P.; Tremblay, Y.D.N.; Jubelin, G.; Jacques, M.; Harel, J. Biofilm-forming abilities of shiga toxin-producing Escherichia coli isolates associated with human infections. Appl. Environ. Microbiol. 2015, 28, 1448–1458. [Google Scholar] [CrossRef] [Green Version]





© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Amaretti, A.; Righini, L.; Candeliere, F.; Musmeci, E.; Bonvicini, F.; Gentilomi, G.A.; Rossi, M.; Raimondi, S. Antibiotic Resistance, Virulence Factors, Phenotyping, and Genotyping of Non-Escherichia coli Enterobacterales from the Gut Microbiota of Healthy Subjects. Int. J. Mol. Sci. 2020, 21, 1847. https://doi.org/10.3390/ijms21051847
Amaretti A, Righini L, Candeliere F, Musmeci E, Bonvicini F, Gentilomi GA, Rossi M, Raimondi S. Antibiotic Resistance, Virulence Factors, Phenotyping, and Genotyping of Non-Escherichia coli Enterobacterales from the Gut Microbiota of Healthy Subjects. International Journal of Molecular Sciences. 2020; 21(5):1847. https://doi.org/10.3390/ijms21051847
Chicago/Turabian StyleAmaretti, Alberto, Lucia Righini, Francesco Candeliere, Eliana Musmeci, Francesca Bonvicini, Giovanna Angela Gentilomi, Maddalena Rossi, and Stefano Raimondi. 2020. "Antibiotic Resistance, Virulence Factors, Phenotyping, and Genotyping of Non-Escherichia coli Enterobacterales from the Gut Microbiota of Healthy Subjects" International Journal of Molecular Sciences 21, no. 5: 1847. https://doi.org/10.3390/ijms21051847
APA StyleAmaretti, A., Righini, L., Candeliere, F., Musmeci, E., Bonvicini, F., Gentilomi, G. A., Rossi, M., & Raimondi, S. (2020). Antibiotic Resistance, Virulence Factors, Phenotyping, and Genotyping of Non-Escherichia coli Enterobacterales from the Gut Microbiota of Healthy Subjects. International Journal of Molecular Sciences, 21(5), 1847. https://doi.org/10.3390/ijms21051847