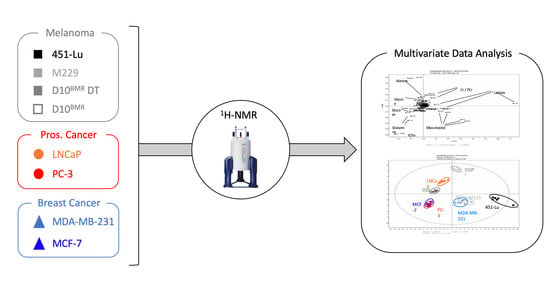
A New Classification Method of Metastatic Cancers Using a 1H-NMR-Based Approach: A Study Case of Melanoma, Breast, and Prostate Cancer Cell Lines
Abstract
:1. Introduction
2. Results
2.1. 1H-NMR Signature from the Cellular Extracts
2.1.1. Identification of Discriminant Metabolites Using Multivariate Data Analysis
2.1.2. Additional Spectra Investigations
2.1.3. Heatmap of the Identified Discriminant Metabolites
2.1.4. Enrichment Analysis of the Intracellular Discriminant Metabolites
2.1.5. Data Transposition to Metabolic Ratios
2.2. Extracellular Compartments Analysis
2.3. Metabolic Inhibition of the Glycolytic, Glutamine, and Choline Pathways
3. Discussion
3.1. 1H-NMR Identification of Metastatic Metabolome Subclasses
3.2. Disrupting Metastatic Metabolome Using Metabolic Inhibitors
4. Materials and Methods
4.1. Cell Lines and Culture
4.2. Metabolic Inhibitors and Viability Test
4.3. 1H-NMR Samples Collection and Extraction
4.4. 1H-NMR Spectroscopy
4.5. Spectra Processing and Multivariate Data Analysis
4.6. Metabolic Signature Validation, Heatmap, and Enrichment
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Liberti, M.V.; Locasale, J.W. The Warburg Effect: How Does it Benefit Cancer Cells? Trends Biochem. Sci. 2016, 41, 211–218. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, L.; Supuran, C.T.; Alfarouk, K.O. The Warburg Effect and the Hallmarks of Cancer. Anticancer Agents Med. Chem. 2017, 17, 164–170. [Google Scholar] [CrossRef] [PubMed]
- Ashton, T.M.; McKenna, W.G.; Kunz-Schughart, L.A.; Higgins, G.S. Oxidative Phosphorylation as an Emerging Target in Cancer Therapy. Clin. Cancer Res. 2018, 24, 2482–2490. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Alesi, G.; Kang, S. Glutaminolysis as a target for cancer therapy. Oncogene 2016, 35, 3619–3625. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Venneti, S.; Nagrath, D. Glutaminolysis: A Hallmark of Cancer Metabolism. Annu. Rev. Biomed. Eng. 2017, 19, 163–194. [Google Scholar] [CrossRef] [PubMed]
- Pavlova, N.N.; Thompson, C.B. The Emerging Hallmarks of Cancer Metabolism. Cell Metab. 2016, 23, 27–47. [Google Scholar] [CrossRef]
- DeBerardinis, R.J.; Chandel, N.S. Fundamentals of cancer metabolism. Sci. Adv. 2016, 2, e1600200. [Google Scholar] [CrossRef]
- Beger, R.D. A Review of Applications of Metabolomics in Cancer. Metabolites 2013, 3, 552–574. [Google Scholar] [CrossRef]
- Wishart, D.S.; Mandal, R.; Stanislaus, A.; Ramirez-Gaona, M. Cancer Metabolomics and the Human Metabolome Database. Metabolites 2016, 6, 10. [Google Scholar] [CrossRef]
- Seyfried, T.N.; Huysentruyt, L.C. On the Origin of Cancer Metastasis. Crit. Rev. Oncog. 2013, 18, 43–73. [Google Scholar] [CrossRef]
- Pascual, G.; Domínguez, D.; Benitah, S.A. The contributions of cancer cell metabolism to metastasis. Dis. Models Mech. 2018, 11, dmm032920. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Hoffmann, A.D.; Liu, H.; Liu, X. Organotropism: New insights into molecular mechanisms of breast cancer metastasis. NPJ Precis. Oncol. 2018, 2, 4. [Google Scholar] [CrossRef] [PubMed]
- Gandaglia, G.; Abdollah, F.; Schiffmann, J.; Trudeau, V.; Shariat, S.F.; Kim, S.P.; Perrotte, P.; Montorsi, F.; Briganti, A.; Trinh, Q.D.; et al. Distribution of metastatic sites in patients with prostate cancer: A population-based analysis. Prostate 2014, 74, 210–216. [Google Scholar] [CrossRef]
- Damsky, W.E.; Rosenbaum, L.E.; Bosenberg, M. Decoding Melanoma Metastasis. Cancers (Basel) 2010, 3, 126–163. [Google Scholar] [CrossRef] [PubMed]
- Dupuy, F.; Tabariès, S.; Andrzejewski, S.; Dong, Z.; Blagih, J.; Annis, M.G.; Omeroglu, A.; Gao, D.; Leung, S.; Amir, E.; et al. PDK1-Dependent Metabolic Reprogramming Dictates Metastatic Potential in Breast Cancer. Cell Metab. 2015, 22, 577–589. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Lee, H.J.; Wu, X.; Huo, L.; Kim, S.J.; Xu, L.; Wang, Y.; He, J.; Bollu, L.R.; Gao, G.; et al. Gain of Glucose-Independent Growth upon Metastasis of Breast Cancer Cells to the Brain. Cancer Res. 2015, 75, 554–565. [Google Scholar] [CrossRef]
- Mashimo, T.; Pichumani, K.; Vemireddy, V.; Hatanpaa, K.J.; Singh, D.K.; Sirasanagandla, S.; Nannepaga, S.; Piccirillo, S.G.; Kovacs, Z.; Foong, C.; et al. Acetate is a bioenergetic substrate for human glioblastoma and brain metastases. Cell 2014, 159, 1603–1614. [Google Scholar] [CrossRef]
- Puchades-Carrasco, L.; Pineda-Lucena, A. Metabolomics Applications in Precision Medicine: An Oncological Perspective. Curr. Top. Med. Chem. 2017, 17, 2740–2751. [Google Scholar] [CrossRef]
- Hirschhaeuser, F.; Sattler, U.G.A.; Mueller-Klieser, W. Lactate: A Metabolic Key Player in Cancer. Cancer Res. 2011, 71, 6921–6925. [Google Scholar] [CrossRef]
- Greenhouse, W.V.; Lehninger, A.L. Occurrence of the malate-aspartate shuttle in various tumor types. Cancer Res. 1976, 36, 1392–1396. [Google Scholar]
- Overview of Cancer Metabolism—Metabolism Research—News Arraystar. Available online: https://www.arraystar.com/reviews/overview-of-cancer-metabolism/ (accessed on 11 April 2019).
- Kumar, S.M.; Yu, H.; Edwards, R.; Chen, L.; Kazianis, S.; Brafford, P.; Acs, G.; Herlyn, M.; Xu, X. Mutant V600E BRAF increases hypoxia inducible factor-1alpha expression in melanoma. Cancer Res. 2007, 67, 3177–3184. [Google Scholar] [CrossRef] [PubMed]
- Shi, Y.; Chang, M.; Wang, F.; Ouyang, X.; Jia, Y.; Du, H. Role and mechanism of hypoxia-inducible factor-1 in cell growth and apoptosis of breast cancer cell line MDA-MB-231. Oncol. Lett. 2010, 1, 657–662. [Google Scholar] [CrossRef]
- Marín-Hernández, A.; Gallardo-Pérez, J.C.; Ralph, S.J.; Rodríguez-Enríquez, S.; Moreno-Sánchez, R. HIF-1alpha modulates energy metabolism in cancer cells by inducing over-expression of specific glycolytic isoforms. Mini-Rev. Med. Chem. 2009, 9, 1084–1101. [Google Scholar] [CrossRef] [PubMed]
- Ullah, M.S.; Davies, A.J.; Halestrap, A.P. The plasma membrane lactate transporter MCT4, but not MCT1, is up-regulated by hypoxia through a HIF-1alpha-dependent mechanism. J. Biol. Chem. 2006, 281, 9030–9037. [Google Scholar] [CrossRef] [PubMed]
- Parmenter, T.J.; Kleinschmidt, M.; Kinross, K.M.; Bond, S.T.; Li, J.; Kaadige, M.R.; Rao, A.; Sheppard, K.E.; Hugo, W.; Pupo, G.M.; et al. Response of BRAF-mutant melanoma to BRAF inhibition is mediated by a network of transcriptional regulators of glycolysis. Cancer Discov. 2014, 4, 423–433. [Google Scholar] [CrossRef]
- Patra, S.; Ghosh, A.; Roy, S.S.; Bera, S.; Das, M.; Talukdar, D.; Ray, S.; Wallimann, T.; Ray, M. A short review on creatine-creatine kinase system in relation to cancer and some experimental results on creatine as adjuvant in cancer therapy. Amino Acids 2012, 42, 2319–2330. [Google Scholar] [CrossRef]
- Yan, Y.B. Creatine kinase in cell cycle regulation and cancer. Amino Acids 2016, 48, 1775–1784. [Google Scholar] [CrossRef]
- Loo, J.M.; Scherl, A.; Nguyen, A.; Man, F.Y.; Weinberg, E.; Zeng, Z.; Saltz, L.; Paty, P.B.; Tavazoie, S.F. Extracellular Metabolic Energetics Can Promote Cancer Progression. Cell 2015, 160, 393–406. [Google Scholar] [CrossRef]
- Bagnoli, M.; Granata, A.; Nicoletti, R.; Krishnamachary, B.; Bhujwalla, Z.M.; Canese, R.; Podo, F.; Canevari, S.; Iorio, E.; Mezzanzanica, D. Choline Metabolism Alteration: A Focus on Ovarian Cancer. Front. Oncol. 2016, 6, 153. [Google Scholar] [CrossRef]
- Glunde, K.; Bhujwalla, Z.M.; Ronen, S.M. Choline metabolism in malignant transformation. Nat. Rev. Cancer 2011, 11, 835–848. [Google Scholar] [CrossRef]
- Glunde, K.; Jacobs, M.A.; Bhujwalla, Z.M. Choline metabolism in cancer: Implications for diagnosis and therapy. Expert Rev. Mol. Diagn. 2006, 6, 821–829. [Google Scholar] [CrossRef] [PubMed]
- Garg, R.; Benedetti, L.G.; Abera, M.B.; Wang, H.; Abba, M.; Kazanietz, M.G. Protein kinase C and cancer: What we know and what we do not. Oncogene 2014, 33, 5225–5237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kolch, W.; Heidecker, G.; Kochs, G.; Hummel, R.; Vahidi, H.; Mischak, H.; Finkenzeller, G.; Marmé, D.; Rapp, U.R. Protein kinase C alpha activates RAF-1 by direct phosphorylation. Nature 1993, 364, 249–252. [Google Scholar] [CrossRef] [PubMed]
- Bunney, T.D.; Katan, M. Phosphoinositide signalling in cancer: Beyond PI3K and PTEN. Nat. Rev. Cancer 2010, 10, 342–352. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Pinedo, C.; El Mjiyad, N.; Ricci, J.E. Cancer metabolism: Current perspectives and future directions. Cell Death Dis. 2012, 3, e248. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Locasale, J.W.; Grassian, A.R.; Melman, T.; Lyssiotis, C.A.; Mattaini, K.R.; Bass, A.J.; Heffron, G.; Metallo, C.M.; Muranen, T.; Sharfi, H.; et al. Phosphoglycerate dehydrogenase diverts glycolytic flux and contributes to oncogenesis. Nat. Genet. 2011, 43, 869–874. [Google Scholar] [CrossRef] [Green Version]
- Baenke, F.; Chaneton, B.; Smith, M.; Van Den Broek, N.; Hogan, K.; Tang, H.; Viros, A.; Martin, M.; Galbraith, L.; Girotti, M.R.; et al. Resistance to BRAF inhibitors induces glutamine dependency in melanoma cells. Mol. Oncol. 2016, 10, 73–84. [Google Scholar] [CrossRef]
- de Queiroz, R.M.; Carvalho, É.; Dias, W.B. O-GlcNAcylation: The Sweet Side of the Cancer. Front. Oncol. 2014, 4, 132. [Google Scholar] [CrossRef] [Green Version]
- Kamigaito, T.; Okaneya, T.; Kawakubo, M.; Shimojo, H.; Nishizawa, O.; Nakayama, J. Overexpression of O-GlcNAc by prostate cancer cells is significantly associated with poor prognosis of patients. Prostate Cancer Prostatic Dis. 2014, 17, 18–22. [Google Scholar] [CrossRef] [Green Version]
- Macheda, M.L.; Rogers, S.; Best, J.D. Molecular and cellular regulation of glucose transporter (GLUT) proteins in cancer. J. Cell. Physiol. 2005, 202, 654–662. [Google Scholar] [CrossRef]
- San-Millán, I.; Brooks, G.A. Reexamining cancer metabolism: Lactate production for carcinogenesis could be the purpose and explanation of the Warburg Effect. Carcinogenesis 2017, 38, 119–133. [Google Scholar] [CrossRef] [PubMed]
- Le, A.; Cooper, C.R.; Gouw, A.M.; Dinavahi, R.; Maitra, A.; Deck, L.M.; Royer, R.E.; Vander Jagt, D.L.; Semenza, G.L.; Dang, C.V. Inhibition of lactate dehydrogenase A induces oxidative stress and inhibits tumor progression. Proc. Natl. Acad. Sci. USA 2010, 107, 2037–2042. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mohamed, A.; Deng, X.; Khuri, F.R.; Owonikoko, T.K. Altered glutamine metabolism and therapeutic opportunities for lung cancer. Clin. Lung Cancer 2014, 15, 7–15. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Márquez, J.; Alonso, F.J.; Matés, J.M.; Segura, J.A.; Martín-Rufián, M.; Campos-Sandoval, J.A. Glutamine Addiction In Gliomas. Neurochem Res. 2017, 42, 1735–1746. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.K.; Islam, S.M.R.; Abdullah-AL-Wadud, M.; Islam, S.; Ali, F.; Park, K.S. Multiomics Analysis Reveals that GLS and GLS2 Differentially Modulate the Clinical Outcomes of Cancer. J. Clin. Med. 2019, 8, 355. [Google Scholar] [CrossRef] [Green Version]
- Ramírez de Molina, A.; Rodríguez-González, A.; Gutiérrez, R.; Martínez-Piñeiro, L.; Sánchez, J.J.; Bonilla, F.; Rosell, R.; Lacal, J.C. Overexpression of choline kinase is a frequent feature in human tumor-derived cell lines and in lung, prostate, and colorectal human cancers. Biochem. Biophys. Res. Commun. 2002, 296, 580–583. [Google Scholar] [CrossRef]
- Hong, B.S.; Allali-Hassani, A.; Tempel, W.; Finerty, P.J.; MacKenzie, F.; Dimov, S.; Vedadi, M.; Park, H.W. Crystal Structures of Human Choline Kinase Isoforms in Complex with Hemicholinium-3. J. Biol. Chem. 2010, 285, 16330–16340. [Google Scholar] [CrossRef] [Green Version]
- Jones, A.T.; Narov, K.; Yang, J.; Sampson, J.R.; Shen, M.H. Efficacy of Dual Inhibition of Glycolysis and Glutaminolysis for Therapy of Renal Lesions in Tsc2 +/− Mice. Neoplasia 2019, 21, 230–238. [Google Scholar] [CrossRef]





© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Schepkens, C.; Dallons, M.; Dehairs, J.; Talebi, A.; Jeandriens, J.; Drossart, L.-M.; Auquier, G.; Tagliatti, V.; Swinnen, J.V.; Colet, J.-M. A New Classification Method of Metastatic Cancers Using a 1H-NMR-Based Approach: A Study Case of Melanoma, Breast, and Prostate Cancer Cell Lines. Metabolites 2019, 9, 281. https://doi.org/10.3390/metabo9110281
Schepkens C, Dallons M, Dehairs J, Talebi A, Jeandriens J, Drossart L-M, Auquier G, Tagliatti V, Swinnen JV, Colet J-M. A New Classification Method of Metastatic Cancers Using a 1H-NMR-Based Approach: A Study Case of Melanoma, Breast, and Prostate Cancer Cell Lines. Metabolites. 2019; 9(11):281. https://doi.org/10.3390/metabo9110281
Chicago/Turabian StyleSchepkens, Corentin, Matthieu Dallons, Jonas Dehairs, Ali Talebi, Jérôme Jeandriens, Lise-Marie Drossart, Guillaume Auquier, Vanessa Tagliatti, Johannes V. Swinnen, and Jean-Marie Colet. 2019. "A New Classification Method of Metastatic Cancers Using a 1H-NMR-Based Approach: A Study Case of Melanoma, Breast, and Prostate Cancer Cell Lines" Metabolites 9, no. 11: 281. https://doi.org/10.3390/metabo9110281
APA StyleSchepkens, C., Dallons, M., Dehairs, J., Talebi, A., Jeandriens, J., Drossart, L. -M., Auquier, G., Tagliatti, V., Swinnen, J. V., & Colet, J. -M. (2019). A New Classification Method of Metastatic Cancers Using a 1H-NMR-Based Approach: A Study Case of Melanoma, Breast, and Prostate Cancer Cell Lines. Metabolites, 9(11), 281. https://doi.org/10.3390/metabo9110281